The root system of Arabidopsis thaliana is an excellent model to study the relationship between cell cycle regulation and growth and development. Understanding this offers a great potential for altering root architecture and water uptake, allowing to design plants to survive under dryer conditions. The research of this group focuses on the formation of lateral root primordia in the pericycle to investigate how cell cycle regulation is involved in the initiation of new organs. Plant roots serve a multitude of functions. They anchor plants and supply them with water and nutrients and exchange various growth substances with the shoots. At the root-soil interface, numerous interactions between plants and their environment take place. The diversity of functions and broad range of interactions with the environment render the biology of roots complicated. During the last ten years, Arabidopsis thaliana has been proven to be an efficient model plant to study root development and time has come to extrapolate the obtained insights to crop species such as maize, a species that is currently also under investigation.
Besides the various advantages this plant offers in genetic and molecular studies, the Arabidopsis root boasts quite predictable ontogeny, very simple anatomy and a high level of transparency, making it very suitable for morphogenetic and cell biology studies. Understanding how root systems develop is crucial for maximizing crop production in a world in which the population is increasing, and the amount of arable land is decreasing. Lateral root formation or root branching is one of the major regulators of root system architecture. During root branching, developmental and environmental controls of cell cycle regulation are crucial. Mitotic activity must be renewed in differentiated pericycle cells that have left the cell cycle. In Arabidopsis, the lateral roots are initiated by local activation of the pericycle cells at the xylem poles (Dhooge et al., 1999). Earlier, a model describing the cell cycle progression preceding the first formative divisions for lateral root initiation was proposed by members of our group (Beeckman et al., 2001). Unique cell cycle control, dissimilar from that in other pericycle cells, was shown to take place in the xylem pericycle. Good knowledge of this tissue-specific cell cycle regulation is therefore needed for a better understanding of lateral root initiation.
To efficiently study the molecular and cytological events during the early stages of pericycle activation, a lateral root-inducible system was developed. Successive treatments with an auxin transport inhibitor and exogenous auxin were used to prevent the first formative divisions and to activate the whole pericycle, respectively. The morphological and molecular results show that, in this inducible system, pericycle activation corresponds to that of spontaneous lateral root initiation, but is enhanced and covers the whole length of the xylem pericycle. In this system, cell cycle progression during the early course of lateral root induction was monitored using histochemical and molecular techniques. The results demonstrated that CDK inhibitory proteins (KRPs) play a previously unknown role in specifically preventing lateral root initiation at the G1-to-S transition (Himanen et al., in preparation). In addition, the analysis of auxin distribution patterns in the system confirmed that auxin is important for the positioning and frequency of the first formative divisions for lateral root initiation (Casimiro et al., 2001).
In Arabidopsis thaliana, Lateral Root Initiation (LRI) is hallmarked by coordinated asymmetric divisions in adjacent xylem pole pericycle cells. Similar to many developmental processes, it is strongly regulated by the plant hormone auxin.
Introduction
A major advantage over other processes is that LRI can be manipulated easily, which allowed us to design a lateral root-inducible system (Himanen et al., 2002). Genome-wide analyses of transcriptional changes upon this synchronized induction of lateral roots resulted in the elucidation of some crucial aspects of LRI (Himanen et al., 2004 – Cell cycle; Vanneste et al., 2005 – SLR/IAA14; De Smet et al., 2008 – ACR4; De Rybel et al., 2010 – GATA23; De Rybel, Audenaert et al. 2012). A tool allowing to combine these datasets into a versatile compendium has been developed (Parizot et al., 2010) and allowed the selection of new candidate genes for the LRI. These datasets were also made available in the eFP browser online tool: LRI eFP browser.
Studying root development at the macro- and micro- evolutionary scale was facilitated the ongoing amelioration of phenotyping techniques, the extent of plant accessions databases and the arise of next-generation sequencing. Two main strategies have been initiated in the Root Development group to investigate micro-evolution of root architecture in Arabidopsis, and macro-evolution of genetic determinants of lateral root development in different species.
Micro-Evolution of Root Architecture in Arabidopsis: Genome-Wide Association Study (GWAS)
Plant sciences has been enriched recently with development of next generation sequencing, adaptation of statistical tools allowing for forward genetic screens in plants and further evolution of molecular tools available for Arabidopsis thaliana. Ultimately those advancements allow to direct plant research into in-depth analysis of pool of gene variability that is especially easily accessible and worldwide representative in Arabidopsis accessions. With data from high-throughput in vitro phenotyping of root architecture traits, genome-wide association studies are conducted. It is a tool used to correlate phenotyping data with SNPs information. As such, we are able to look more closely into what genes are underlying the conserved underground phenotypes in chosen A. thaliana accessions.
We additionally evaluate the variability and stability of the traits in our in soil, rhizotron systems. This creates an opportunity to broaden the insight into certain underground phenotypes and even link it with shoot development and overall plant’s performance up to its flowering stage.
Macro-Evolution of Lateral Root Development
Most of the knowledge about root development has been acquired from work in the model plant Arabidopsis. Although the regulatory mechanisms of lateral root initiation and root systems architecture are to a large extent unravelled, it is not known how universal this is or in what extent gene functions were conserved through the evolution. Therefore, in addition to the study of lateral root initiation in crops, we are studying lateral root initiation and root branching in several, evolutionary distinct, model plants. Used models include the early land plant and lycophyte Selaginella moellendorffii, the fern Ceratopteris richardii (C-fern) and angiosperms such as Medicago truncatula and Oryza sativa (rice).
Roots of Selaginella originated independently from roots of higher plants. They do not branch by lateral root formation, but by dichotomously splicing of the root tip. Despite of the independent evolution, some root development mechanisms from higher plants seems to be adopted by lycophytes too. Our research focusses on the commonly adopted regulators, as such revealing essential pathways in root development.
In ferns, root develops differentially compared to angiosperms. For example, lateral root induction is not affected by auxins or auxin transport inhibitors (as applied in the Lateral Root Inducible System in Arabidopsis).
LRI datasets were compiled and integrated in the eFP browser online tool. Genome-wide analyses of transcriptional changes upon this synchronized induction of lateral roots resulted in the elucidation of some crucial aspects of LRI (Himanen et al., 2004 – Cell cycle; Vanneste et al., 2005 – SLR/IAA14; De Smet et al., 2008 – ACR4; De Rybel et al., 2010 – GATA23; De Rybel, Audenaert et al. 2012). A tool allowing to combine these datasets into a versatile compendium has been developed (Parizot et al., 2010) and allowed the selection of new candidate genes for the LRI. These datasets were also made available in the eFP browser online tool: LRI eFP browser.
LRI datasets compendium
Here is a snapshot of the relative expression of GATA23 in these experiments (low expression values are masked in grey).
Within the angiosperms, major differences in root architecture between dicotyledonous and monocotyledonous plants exist. We are working on the genetic and molecular mechanisms involved in the development of the root system in major crop species.
Introduction
Because of the different roles the root system plays in overall plant growth, root architecture is a fundamental aspect of plant growth and development. The root system especially acquires water and nutrients from the soil, anchors the plant in the substrate, synthesizes hormones and metabolites, interacts with symbiotic microorganisms, and insures storage functions. In light of these characteristics, more and more breeders focus their attention on this underground organ in order to increase yield. This requires a better understanding of the relation of this part of the plant with the environment and of its highly adaptive behavior (Lynch, 2007; Gewin, 2010; Den Herder et al., 2010). Within the angiosperms, major differences in root architecture between dicotyledonous and monocotyledonous plants exist. Dicots develop a tap root system composed of a main primary root, already formed during embryogenesis, which grows vertically into the soil and gives rise to the emergence of numerous lateral roots extending the surface area. Monocots have a fibrous root system in which the embryonic primary root is only important for the early development of the plant (Feix et al., 2002) and in which an extensive postembryonic shoot-born root system is formed later on. Very little is known about the genetic and molecular mechanisms involved in the development and architecture of the root system in major crop species, generally monocotyledonous plants. Lack of insight is certainly a consequence of the difficulty to access and observe this organ in its natural habitat, namely the soil. Moreover, and probably because of this hidden character, the root has been neglected for a long time in crop improvement and in agricultural approaches aiming at increasing shoot biomass. Nevertheless, while most of the work has been done on Arabidopsis thaliana, the awareness of the importance of the root system in modulating plant growth, together with progress in sequencing and new molecular techniques, has caused renewed interest in understanding molecular mechanisms in crop species (Hochholdinger and Zimmermann, 2008; Coudert et al., 2010; Parizot et al., 2012).
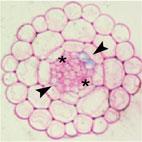
In Arabidopsis thaliana, lateral-root-forming competence of pericycle cells is associated with their position at the xylem poles and depends on the establishment of protoxylem-localized auxin response maxima. In maize, our histological analyses revealed an interruption of the pericycle at the xylem poles, and confirmed the earlier reported proto-phloem-specific lateral root initiation. Phloempole pericycle cells were larger and had thinner cell walls compared with the other pericycle cells, highlighting the heterogeneous character of the maize root pericycle. A maize DR5::RFP marker line demonstrated the presence of auxin response maxima in differentiating xylem cells at the root tip and in cells surrounding the proto-phloem vessels. Chemical inhibition of auxin transport indicated that the establishment of the phloem-localized auxin response maxima is crucial for lateral root formation in maize, because in their absence, random divisions of pericycle and endodermis cells occurred, not resulting in organogenesis. These data hint at an evolutionarily conserved mechanism, in which the establishment of vascular auxin response maxima is required to trigger cells in the flanking outer tissue layer for lateral root initiation. It further indicates that lateral root initiation is not dependent on cellular specification or differentiation of the type of vascular tissue.