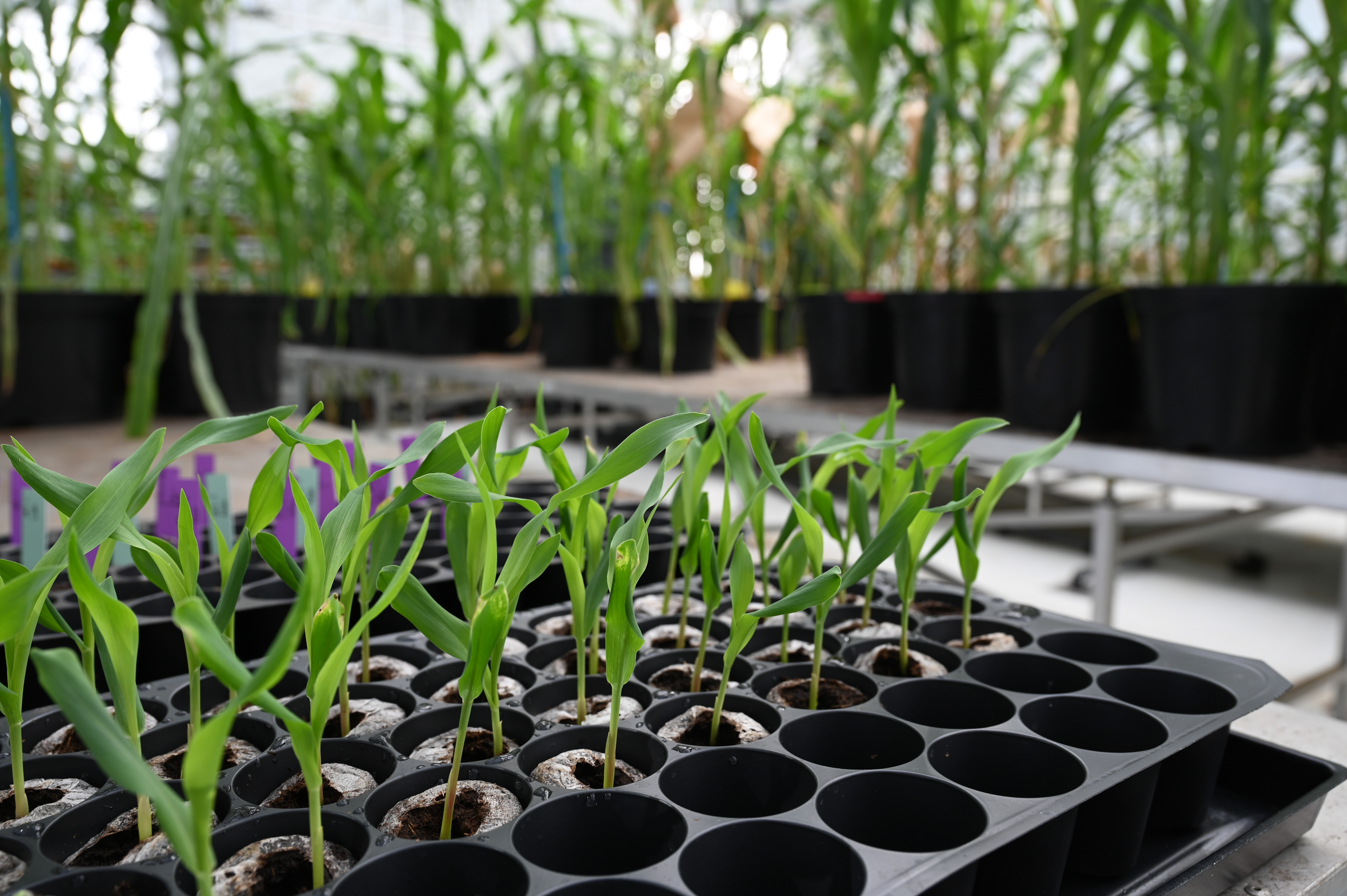

Plant metabolism, growth and development
Many research lines at the Center aim to unravel the genetic programs that control plant metabolism, growth and development. Ultimately, these programs determine the morphological and biochemical features of plants that enable them to optimize carbon fixation, adapt to their environment, and make them suitable for human consumption and utilization.
This focus area on plant metabolism, growth and development has considerable potential to address climate-related issues. Engineering of metabolic pathways can optimize nitrogen use efficiency to reduce greenhouse gas emissions, and increase plant resilience to pathogens and environmental stresses including heat, cold, drought, or heavy metals. Metabolic engineering as well as targeted modification of cell proliferation can optimize carbon capture and support the switch to a bio-based economy. Modulating plant architecture via gene regulatory networks or hormone signaling also can increase carbon sequestration and contribute to improved plant performance under both favorable and environmental stress conditions. Understanding the role of cellular processes such as autophagy, endocytosis, nitrogen and phosphate signaling, or programmed cell death can identify alternative means to increase plant performance.
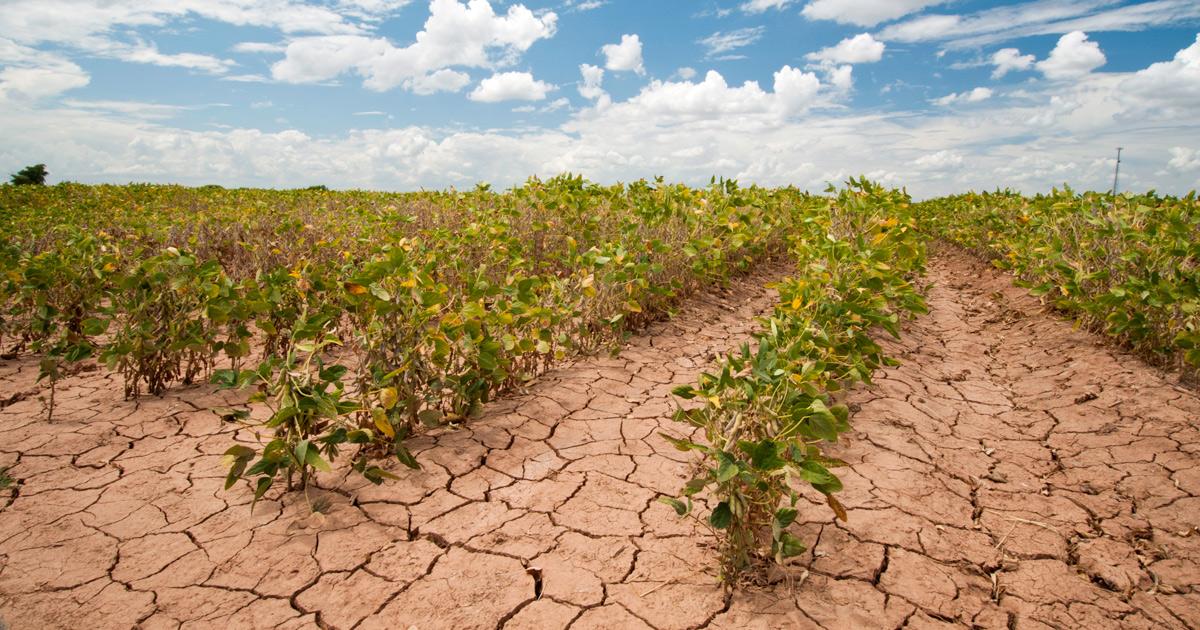

Interaction of plants with the environment
Crop productivity is the most vulnerable factor under climate change. Diverse abiotic stresses such as drought, flooding, waterlogging, high salinity, extreme temperatures, nutrient deficiency and heavy metals reduce crop yield. In this context, researchers at PSB are focusing on the precise elucidation of physiological and molecular mechanisms that contribute to stress tolerance in plants at different developmental stages and in different organs including leaves, roots as well as the soil microbiome that interacts with roots.
An important future aspect of this focus area is the study of the interaction of different abiotic stress factors. Whereas the interaction of some stress factors, e.g. drought and heat, are well documented, others are much less. For example, it is of interest to study how water deficiency affects nutrient uptake by roots and the associated microbiome. This focus area will also examine the relationship between data/phenotypes obtained in the lab compared to similar data obtained from field grown plants. The ultimate aim is to close this lab-field gap as much as possible. Another interesting aspect is intercropping in which different plant species are grown together resulting in higher yields and a better protection to environmental stress. PSB will further explore collaborations to develop and test equipment for measuring the effects of stress on overall plant physiology. The phenotyping platforms will be updated and new, custom-made phenotyping platforms will be installed. For example, an automated imaging system (rhizotron) for the analysis of root growth and architecture is envisaged. Results of this focus area are not only of importance for understanding the molecular basis of stress responses in plants but they can also be used as a basis to develop new strategies for improving sustainability and climate resilience of crop productivity.
Bioinformatics and integrative biology
Bioinformatics and computational biology are cornerstones of modern-day molecular biology and are strongholds of the center. PSB has a longstanding expertise in many subdomains of bioinformatics, including the assembly and annotation of novel plant genome sequences, evolutionary genomics, the analysis and integration of -omics data, gene network modeling and comparative genomics.
While many projects at PSB are already collaborations between experimental and bioinformatics groups, a further integration of dry- and wet-lab activities remains a key goal of the Center. Such further integration is a prerequisite for running complex integrative biology projects on climate change involving e.g. field trials, massive -omics data generation with single cell resolution, phenotyping, data integration for gene function inference, model building for trait prediction, genome editing of candidate genes and validation experiments. Next to efforts to enhance dry-lab skills in wet-labs and vice versa, we strive to increase in the number of joint projects between dry-lab and wet-lab groups tackling common research interests, and stimulate such bottom-up initiatives where possible.
Genome editing will be pivotal for future crop improvement but when dealing with complex biological problems one of the most difficult tasks is to prioritize the list of candidate genes to target. Although great progress has already been made in Arabidopsis, translating this knowledge to crops is hampered by evolutionary divergence between plant species. In particular, the omnipresence of polyploidy in plants hampers both direct translation of knowledge from models to crops and prioritization of candidate genes inferred from -omics analyses on the crop of interest. The bioinformatics groups at PSB have ample expertise in polyploidy, ancient genome duplications, evolutionary genomics and comparative genomics and will continue to both generate fundamental insights and develop tools in these areas. This expertise will be crucial in PSB efforts to link genes to traits in maize, wheat, poplar and soybean, and for investigating how plants cope with climate change.
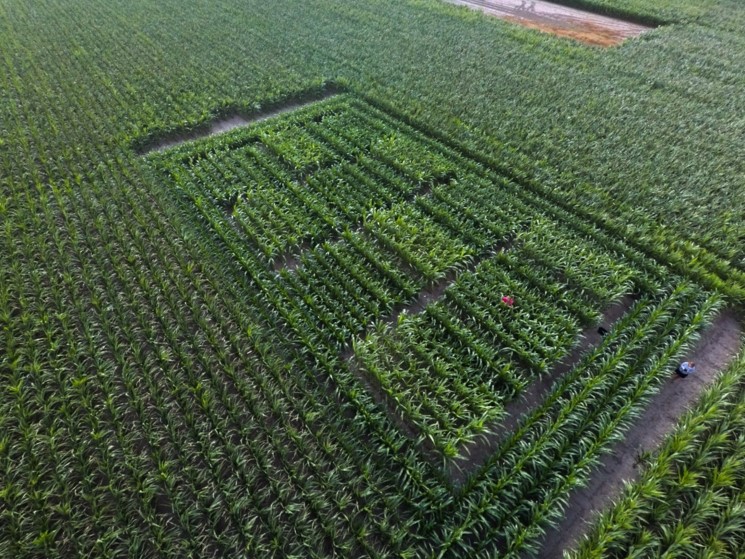

Translation from models to crops and from lab to field
Our ambition is to accelerate the innovation cycle between lab and field. Furthermore, we will strengthen activities to understand how molecular pathways behave in field conditions in order to bring this information back to the lab. Closing the lab-to-field-to-lab learning cycle will be pivotal in translating basic research findings to field applications.
PSB has also developed unique ways to translate basic scientific insights into field applications. Maize and poplar have since long been used in PSB to validate novel biological insight even up to the level of field trials, meanwhile, PSB also initiated research on wheat. Currently, PSB has ongoing projects in wheat on phosphoproteomics, single cell analysis, multiplex mRNA in situ hybridization, genome editing and network biology. In 2021, we will also make wheat transformation available for PSB. Furthermore, we envisage that soybean will be added to the translational pipeline. Soybean is one of the major biotech crops and there is rapidly growing interest to develop locally-adapted soybean as a major European protein crop.
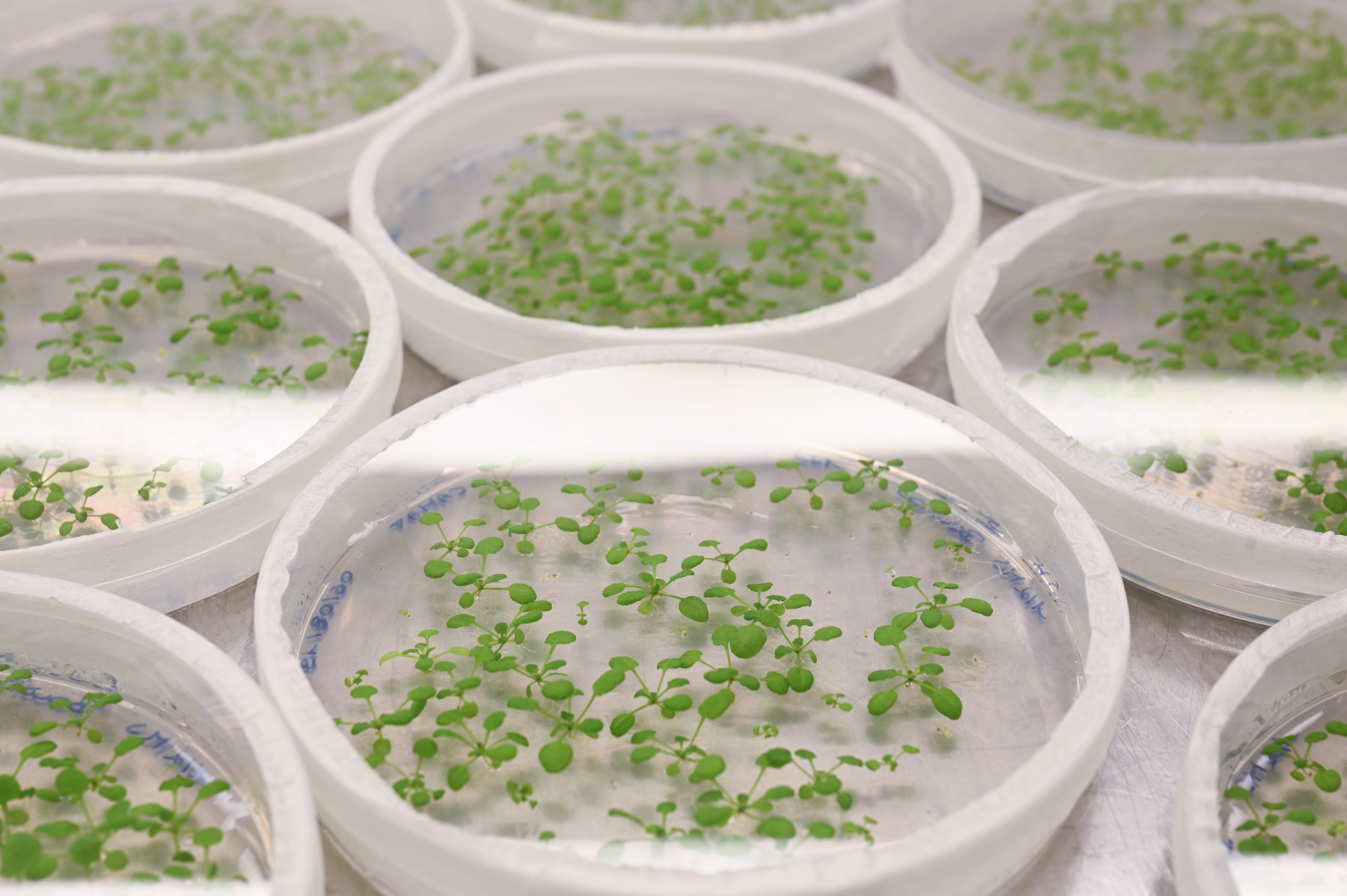

Breakthrough technologies
Breakthrough technologies are the foundation of scientific advances in both basic and translational research and PSB continues to invest and explore in these areas. During the last years, major efforts were made to uncover transcriptional information at high resolution using single-cell technologies. This capability will now be coupled to genome editing and efficient transformation pipelines to improve plant growth, yield, and climate resilience.
To enable the next generation of gene editing methods in crops and other plants, PSB will pursue rapid implementation of genotype-independent transformation, gene editing (including base editing, combinatorial CRISPR screens, and prime editing) and push towards the implementation of more ‘DNA-free’ gene editing methods. Being at the forefront of plant transformation will lower the threshold for our researchers to perform basic research directly in crops and will facilitate translating important findings to relevant varieties. Single cell technologies and spatial transcriptomics will be central in refining molecular networks. Spatial transcriptomics allows in situ expression analysis of hundreds of genes simultaneously and will thus be of specific interest in crop species, which typically lack vast collections of reporter lines and have slower transformation cycles. Furthermore, we explore the use of multiplex gene perturbations in single cells (e.g. protoplasts) to uncover the structure and behavior of transcriptional networks, or the use of a pooled combinatorial CRISPR library approach in cell suspension cultures to select, for instance, combinatorial editing events that optimize cell growth and proliferation under a specific abiotic stress.